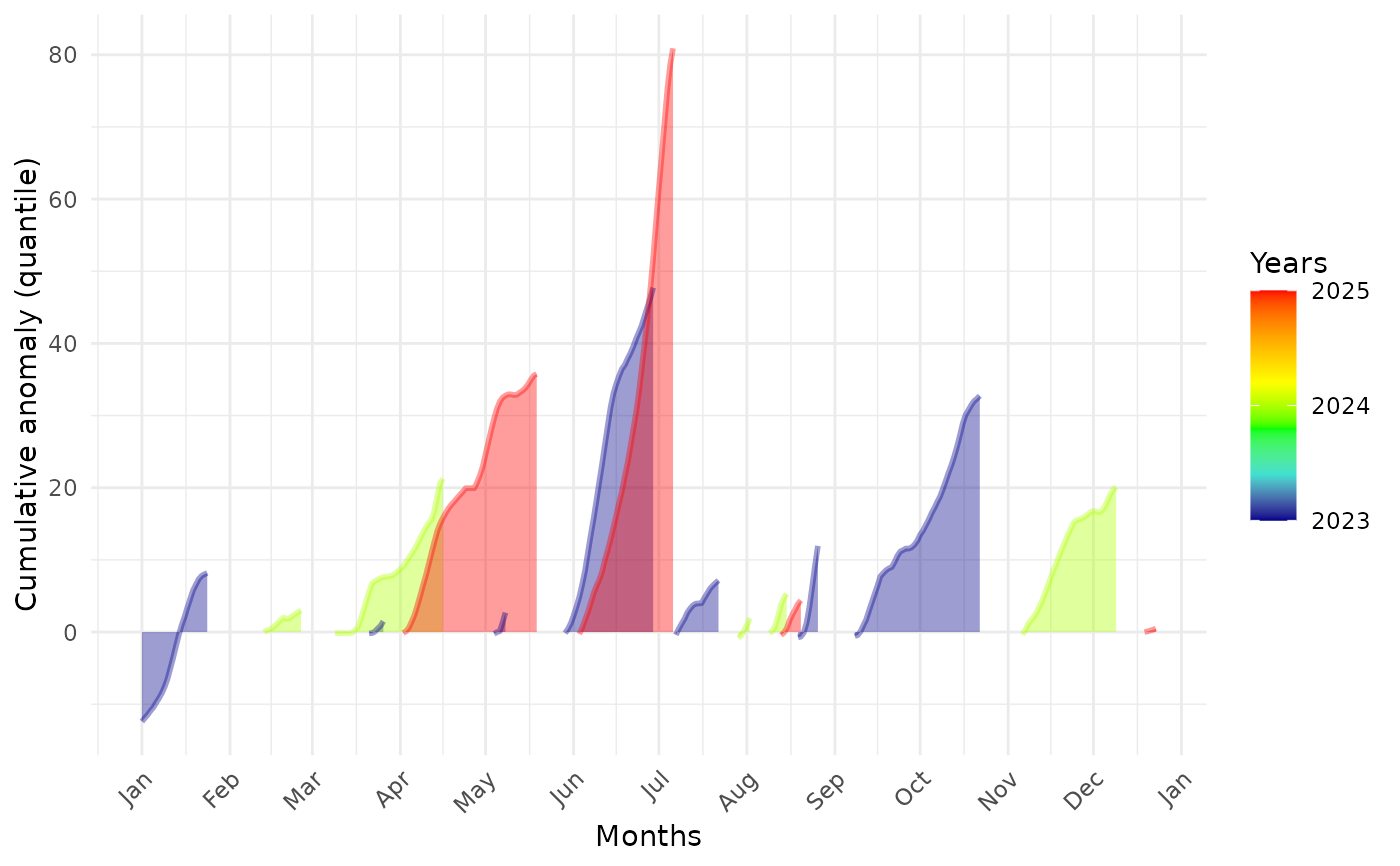
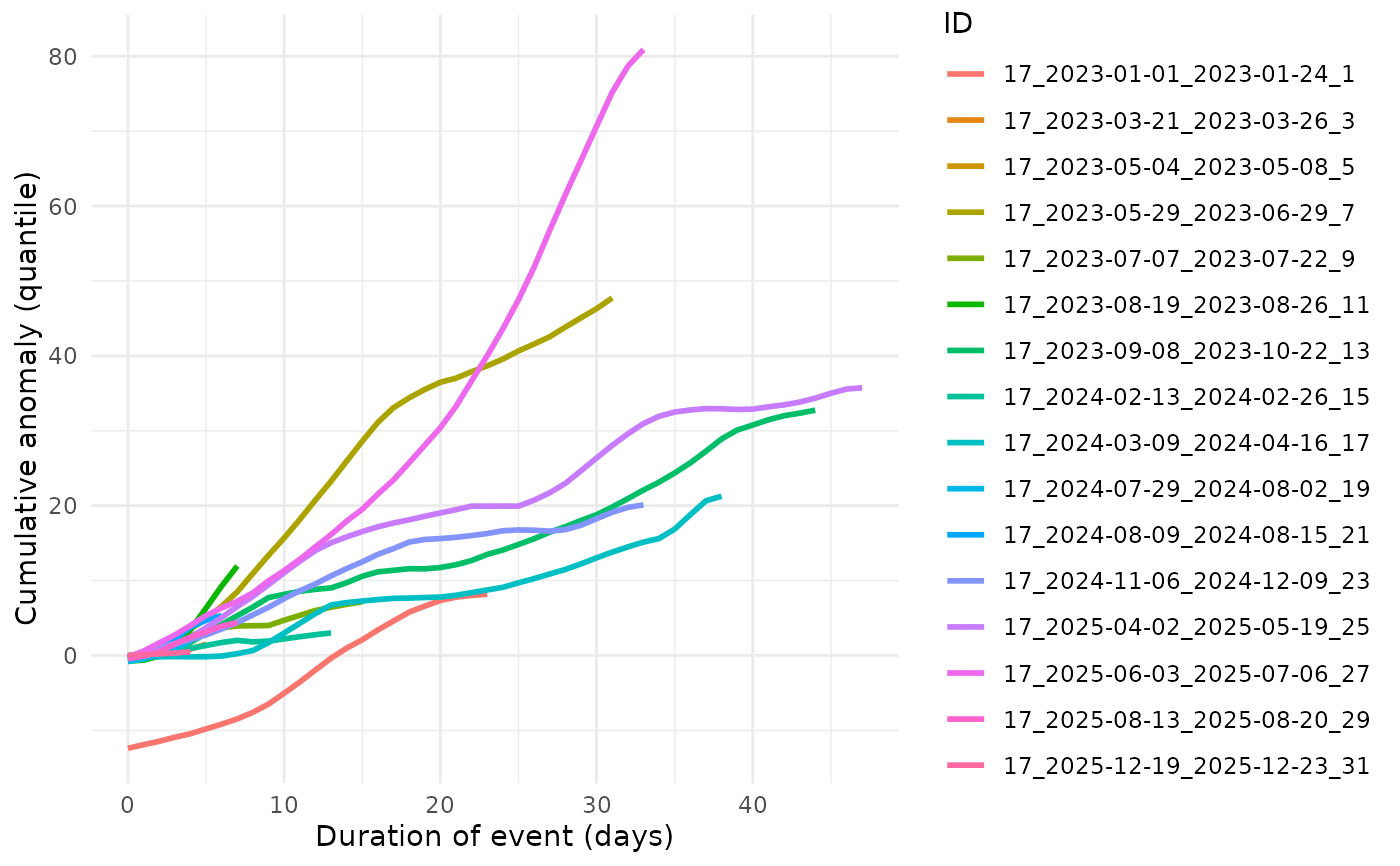
Cumulative anomaly against months of the years
Source:R/bee_plot_cum_ano_season.R
BEE.plot.cumulative_anomaly.RdAllows to see at which time of the year a place is explosed to the bigest anomalies.
BEE.plot.cumulative_anomaly(
metric_point_df,
start_date = NULL,
end_date = NULL,
...
)Arguments
- metric_point_df
: The output of BEE.calc.metric_point() for a given location. Please note that the output of BEE.calc.metric_point() is a list. You only need to provide a dataframe from that list, not the whole list.
- start_date
: = NULL by default or the date at which you want to start the plot. It must be in the same format than metric_point_df$date.
- end_date
: = NULL by default or the date at which you want to stop the plot. It must be in the same format than metric_point_df$date.
- ...
: Customising the graph is possible by adding any general ggplot2 argument.
Value
A 'ggplot2' with the months on the x-axis, the cumulative anomaly per extreme event of the studied parameter value on the y-axis.
Examples
# Load data:
file_name_1 <- system.file(file.path("extdata", "metrics_points_pixel.rds"),
package = "BioExtremeEvent")
metrics_points_day <- readRDS(file_name_1)
# Get data set for one gps point
metrics_points_day <- metrics_points_day[[1]] # first gps point
BEE.plot.cumulative_anomaly(metric_point_df = metrics_points_day,
start_date = NULL,
end_date = NULL)